This function extracts contours from a neuroimaging NIFTI file by identifying
the boundary of the non-zero image support in a selected z-slice.
These contours serve as input for Triangulation::TriMesh,
which is used in Simultaneous Confidence Corridors (SCCs) calculations.
It is highly recommended that the input NIFTI file be pre-processed such that zero values represent the background and non-zero values represent regions of interest.
Arguments
- niftiFile
character, the path to the NIFTI file containing neuroimaging data. Ideally, the file should be masked so that zero values represent the background.- paramZ
integer, the specific z-slice to extract contours from. Default is35.- plotResult
logical, ifTRUE, plots the extracted contours. Default isFALSE.
Value
A list of data frames, where each data frame contains the x and y coordinates of a contour.
The first element typically represents the external boundary, while subsequent elements (if present)
represent internal contours or holes. Each data frame has two columns:
x- x-coordinates of the contour points.y- y-coordinates of the contour points.
Details
This function extracts contours from a NIFTI file, typically a masked image where background values are zero and regions of interest contain non-zero values. Contours are computed from the boundary of the non-zero support of the selected slice.
The extracted contours are typically used as input to Triangulation::TriMesh
to create a triangular mesh of the region, which is then used for
Simultaneous Confidence Corridors calculations.
See also
Triangulation::TriMesh for the next step in the SCC calculation process.
Examples
# Get the file path for a sample NIfTI file
niftiFile <- system.file("extdata", "syntheticControl1.nii.gz", package = "neuroSCC")
# Extract contours from the non-zero support
contours <- neuroContour(niftiFile, paramZ = 35, plotResult = TRUE)
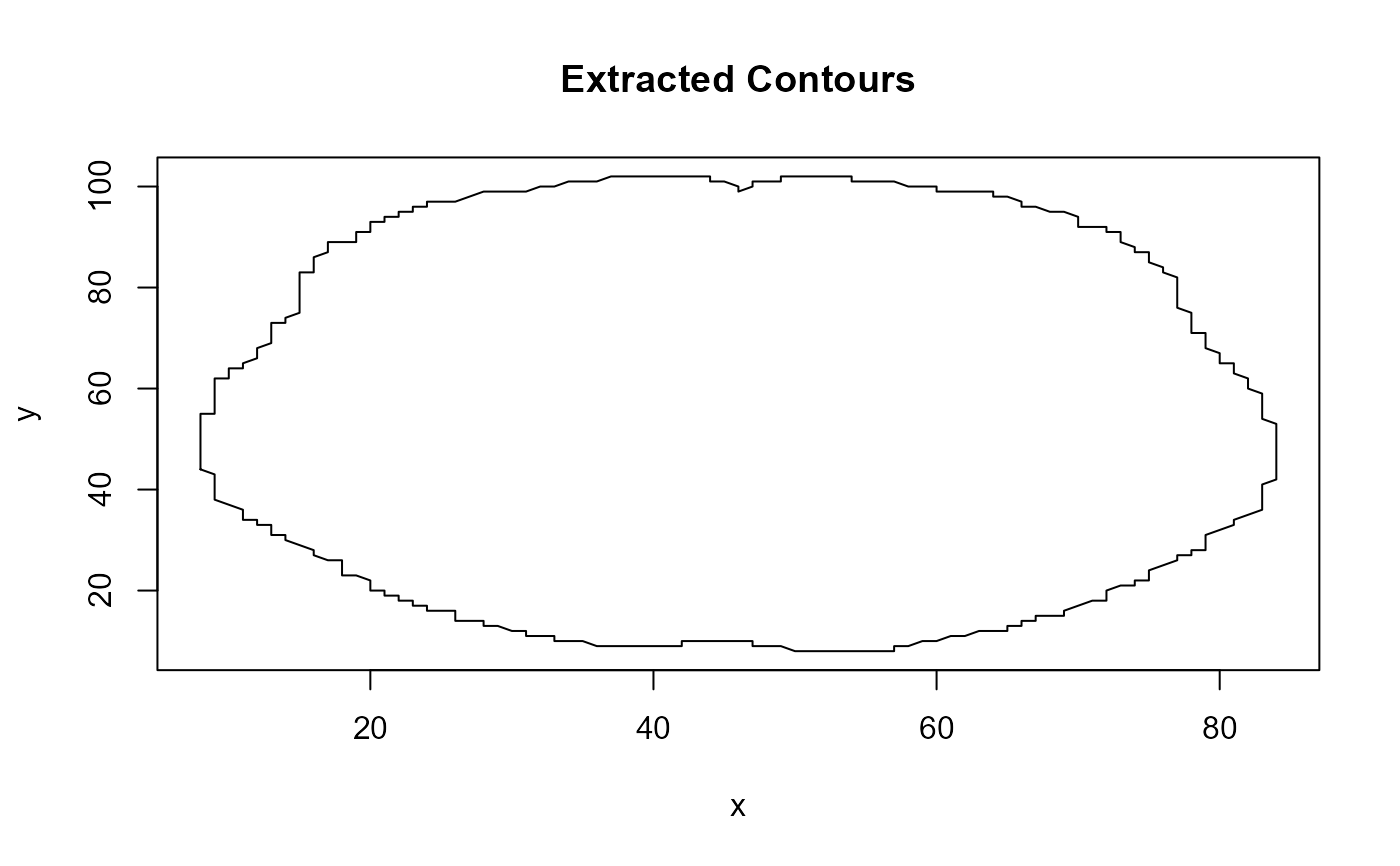 # Display the first few points of the main contour
head(contours[[1]])
#> x y
#> 1 9.0 43.1
#> 2 8.1 44.0
#> 3 8.1 45.0
#> 4 8.1 46.0
#> 5 8.1 47.0
#> 6 8.1 48.0
# Display the first few points of the main contour
head(contours[[1]])
#> x y
#> 1 9.0 43.1
#> 2 8.1 44.0
#> 3 8.1 45.0
#> 4 8.1 46.0
#> 5 8.1 47.0
#> 6 8.1 48.0
